SATIR
Precision Patient–Trial Matching
Scalable High-Recall Constraint-Satisfaction-Based
Information Retrieval for Clinical Trials Matching
LLMs compile criteria into formal constraints.
Relational algebra retrieves. SMT verifies.
1Stanford 2UCLA 3Emory 4Mayo Clinic
*Equal contribution
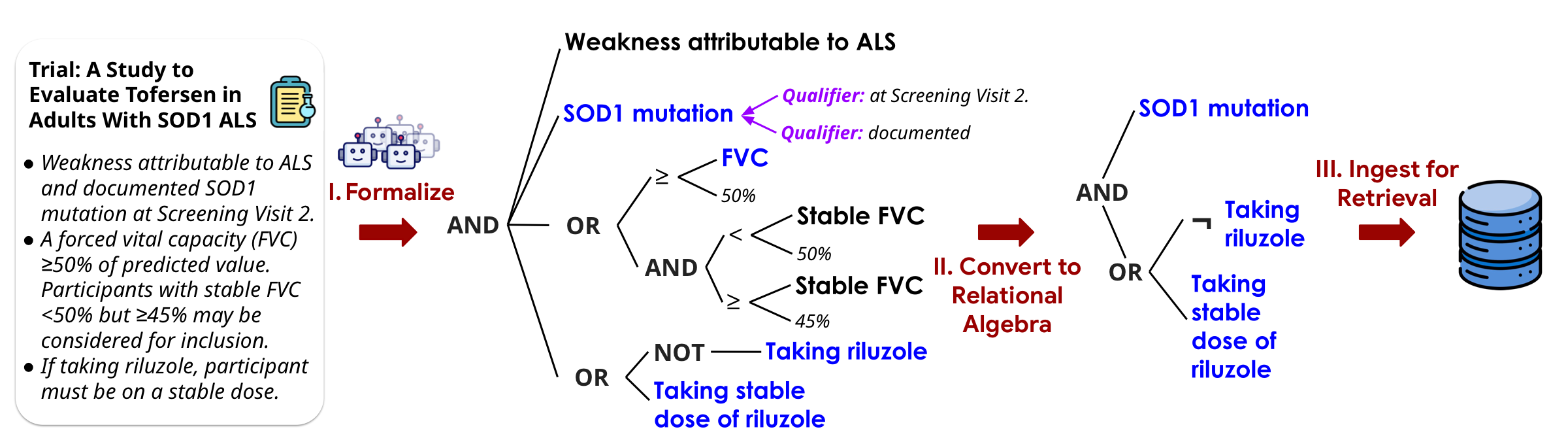
Today's search uses keywords and embedding similarity — soft scores that break down when queries carry hard requirements. A patient either meets an eligibility criterion or doesn't. These are logical predicates, not fuzzy matches.
Encode requirements as formal, composable predicates and solve them exactly using SMT + relational algebra. Use LLMs to convert informal reasoning into explicit, precise, controllable formal constraints.
Nothing is missed
Every decision is traceable
Fix once, propagates everywhere
Evaluated on 59 patients and 3,621 trials, SATIR outperforms TrialGPT on all three retrieval objectives — retrieving 32%–72% more relevant-and-eligible trials per patient, improving recall by 22–38 points, in just 0.15 seconds per patient.
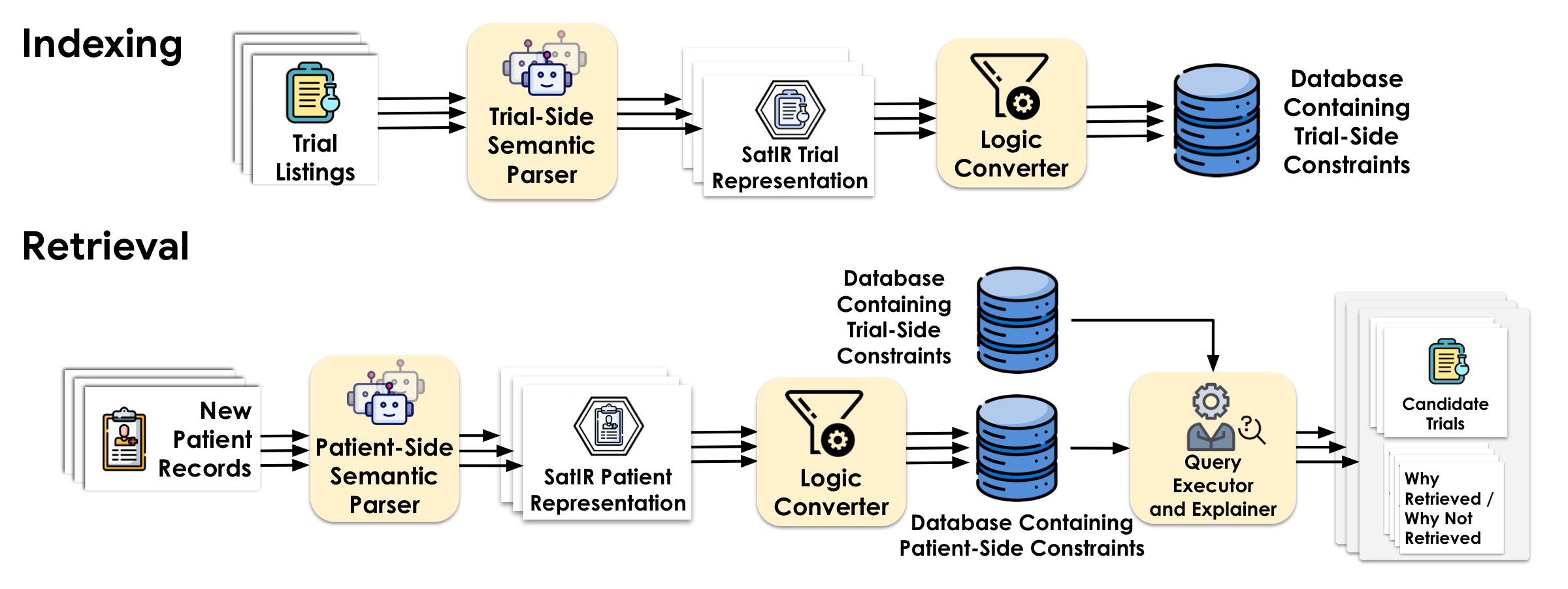
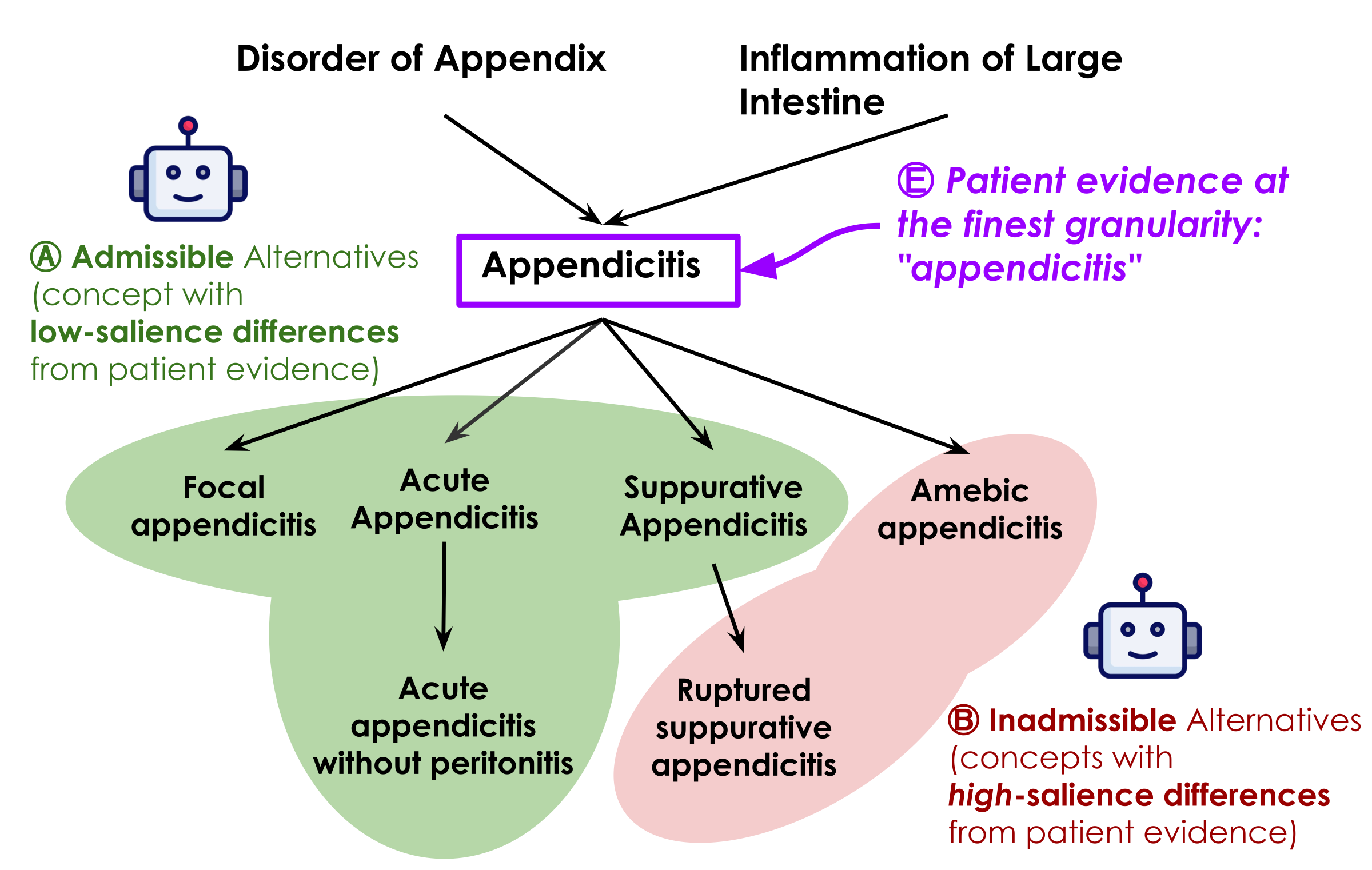
Ontology-Grounded Representation
An SMT-based representation grounded in SNOMED CT captures Boolean logic, numeric thresholds, temporal windows, and ontology-aware concept identity and entailment.
Accurate Semantic Parsing
LLM-based parsers translate complex clinical text — handling underspecified criteria, negation, temporality, and implicit assumptions — into executable formal constraints.
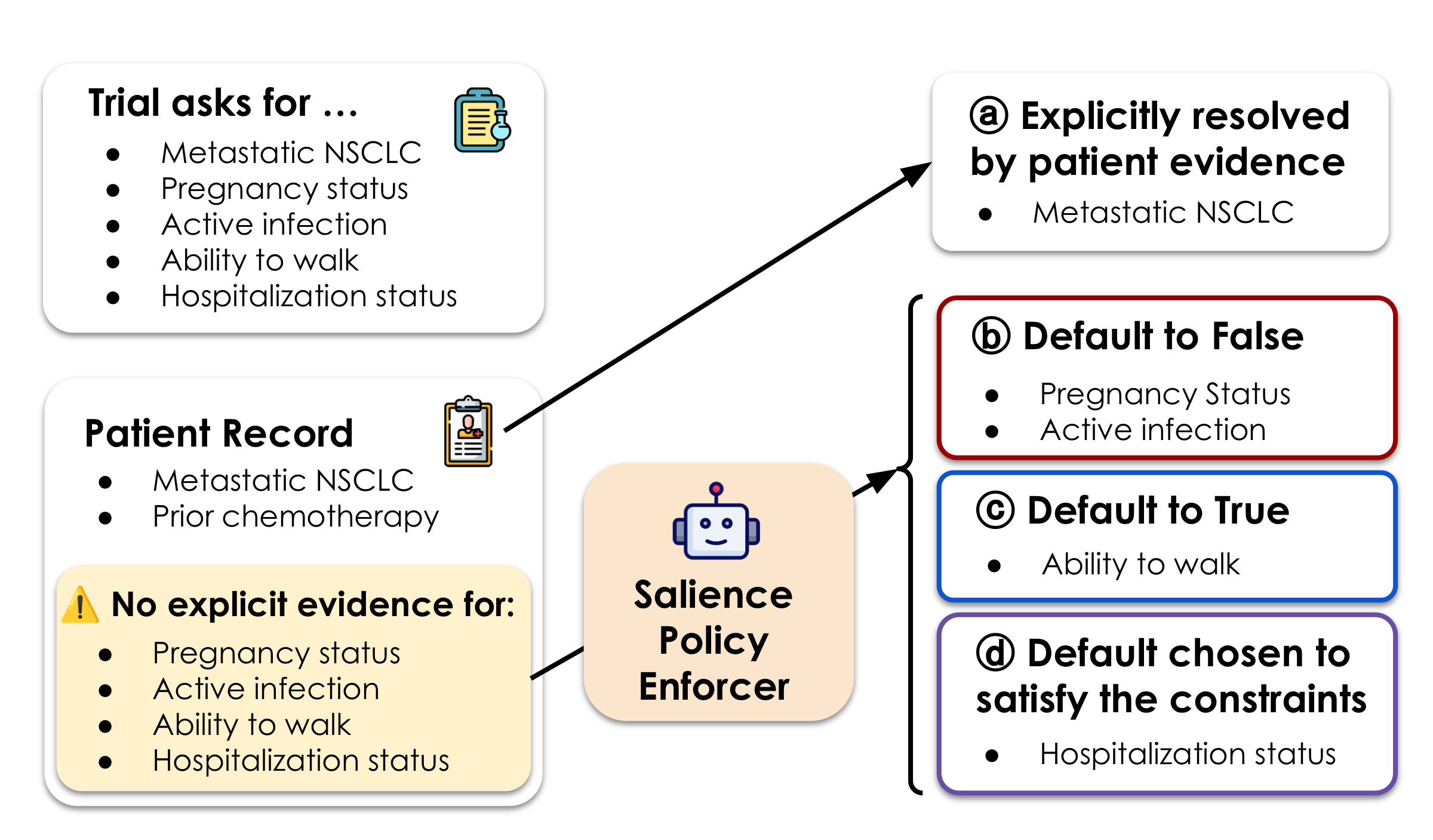
Salience-Based Missingness
SATIR introduces salience to resolve ambiguity from incomplete records — using LLMs to assess whether missing data is clinically significant enough to affect eligibility, with explicit auditable judgments.
Scalable Database Retrieval
SMT constraints are projected into relational algebra and stored in SQL, enabling fast recall-preserving retrieval across hundreds of thousands of trials in under 3 seconds.
Constraint Semantic Parsing
LLM-based pipelines that translate free-text trial criteria and patient records into formal SMT constraint programs.
→ ExploreConstraint Augmentation
Enriching constraints with ontology-grounded subsumption and salience-based handling of incomplete patient records.
→ ExploreConstraint Indexing
Projecting SMT constraints into recall-safe relational algebra and storing them as first-class data in a SQL database.
→ ExploreConstraint Execution
Executing patient constraints against the pre-built trial database via SQL queries in ~0.15 seconds per patient.
→ ExploreRelational Algebra & SMT
SMT (Satisfiability Modulo Theories) decides whether logical formulas over rich types can be satisfied. Relational algebra is the math behind SQL — fast and scalable. SATIR uses relational algebra wherever possible, falling back to SMT only when needed.
“Patient is at least 18 and has a diabetes diagnosis.”
Simple filters — milliseconds over millions of records.
“Eligible iff there exists an assignment of unresolved facts (menopausal status, washout completion) satisfying all constraints, with no assignment implying pregnancy risk.”
Existential + universal reasoning — true constraint solving.
Constraints as First-Class Objects
Constraints are programmable objects — not opaque weights. Each criterion can be composed, inspected, and manipulated: capturing qualifier relationships, expressing complex logic, enforcing consistency, and injecting domain ontologies.
See SMT programming examples →
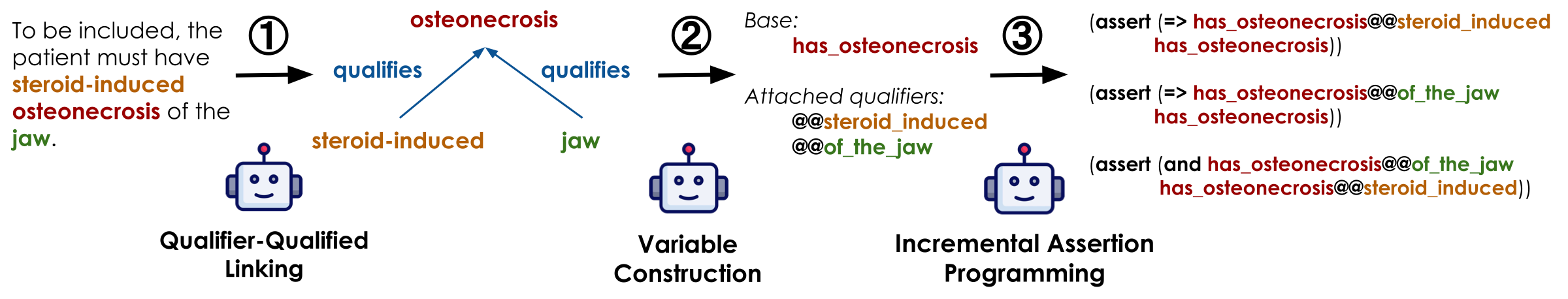
Once constraints are compiled as first-class objects, they can be stored, indexed, and executed directly against a database. Retrieval proceeds in three stages: compatible atom matching, clause-level support, and all-clause aggregation. A patient-trial pair is returned only when every retrieval-relevant constraint is satisfied.
Explore the full execution pipeline →
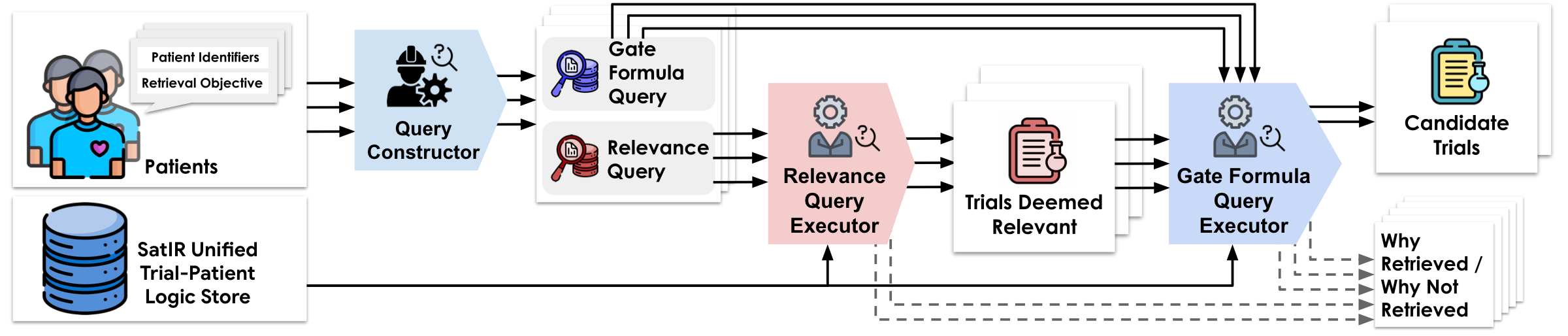
The Boundary of Formalizability
Not everything can be formalized — and it doesn't need to be. SATIR finds the thin boundary: recurring structures get formalized; everything else is gracefully dropped to preserve recall.
Explore subsumption & ontology details →
SNOMED CT, ICD-10, etc. — precise hierarchies, synonyms, and semantic relationships. The most readily usable model.
For domains without an ontology — discover semantic neighborhoods, near-synonyms, and hierarchies on the fly.
Handling Missingness with Salience
Records are routinely incomplete. If a fact is missing, is it because it doesn't apply — or because no one wrote it down? SATIR introduces salience: if a clinically important fact would always be recorded when present, its absence is evidence. Unlike end-to-end LLM matchers, these judgments are explicit, auditable, and overridable.
| Objective | Retrieved/Patient | BMRetriever | ClinBest | BGE-large | PubMedBERT | TrialGPT | SATIR |
|---|---|---|---|---|---|---|---|
| Treat-Chief | 36 | 1.44 | 1.75 | 2.07 | 2.00 | 2.17 | 3.25 |
| Treat-Any | 64 | 2.00 | 2.54 | 2.85 | 2.68 | 2.98 | 5.12 |
| Relevant-to-Any | 121 | 6.39 | 7.24 | 8.68 | 8.92 | 8.93 | 11.76 |
| Objective | SATIR wins | Tie | TrialGPT wins | Served by SATIR | Served by TrialGPT |
|---|---|---|---|---|---|
| Treat-Chief | 20 | 33 | 6 | 39 | 35 |
| Treat-Any | 29 | 28 | 2 | 42 | 39 |
| Relevant-to-Any | 35 | 17 | 7 | 54 | 50 |
Interpretable, auditable, and permanently correctable. Because every eligibility decision traces back to explicit symbolic constraints — not opaque model weights — failures are diagnosable. When a parsing error or an ontology gap is identified and fixed, that fix propagates immediately and permanently across every patient and every trial that touches the corrected predicate. There is no need to retrain, re-embed, or re-evaluate a model. This is the core advantage of SATIR's formal approach: errors are local, fixes are global.
@article{zhou2026satir, title = {Scalable High-Recall Constraint-Satisfaction-Based Information Retrieval for Clinical Trials Matching}, author = {Cyrus Zhou and Yufei Jin and Yilin Xu and Yu-Chiang Wang and Chieh-Ju Chao and Monica S. Lam}, note = {Under review}, year = {2026}, }
Clinical trials matching is just the beginning. The core insight behind SATIR — that real-world search is fundamentally about satisfying structured constraints, not matching keywords — unlocks a new paradigm for information retrieval across high-stakes domains. Anywhere people struggle with complex, multi-faceted requirements, SATIR's formal approach can deliver exhaustive, interpretable, and auditable results.
Go beyond keyword matching — formally verify candidate-role fit across skills, certifications, experience levels, and visa requirements.
Recommend treatments that provably satisfy a patient's full clinical profile — history, contraindications, drug interactions, and guideline criteria.
Plan trips that simultaneously honor budgets, date windows, dietary needs, accessibility, and personal preferences — no compromises, no missed options.
Surface every relevant statute, regulation, or precedent that applies to a specific jurisdiction, timeline, and subject — with full traceability.
Find products that truly meet every specification — compatibility, dimensions, certifications, and performance thresholds — with zero false positives.
Let's build the future of search together. We are actively seeking collaborators to bring constraint-based retrieval to new domains. Whether you have a dataset, a domain challenge, or a vision — we'd love to hear from you.
We gratefully acknowledge support from the Verdant Foundation, the Hasso Plattner Institute, Microsoft Azure AI credits, Itaú Unibanco, BMO Financial Group, and the Stanford HAI Institute. Cyrus Zhou is partially supported by the Stanford School of Engineering Fellowship.
We thank Shengguang Wu for helping get this project started, Yucheng Jiang for the clinician annotation interface, Ning Tang from CreditEase, and our collaborators at Stanford and Mayo Clinic for their expert guidance.








